Cellular Heterogeneity
Clonal microbial populations often harbor rare phenotypic variants that are typically hidden within the majority of the remaining cells, but are crucial for the population’s resilience to external perturbations (Lidstrom and Konopka 2010, Nature Chemical Biology 6: 705). Persister and viable but non-culturable (VBNC) cells are two important clonal bacterial subpopulations that can survive antibiotic treatment. Both persister and VBNC cells pose a serious threat to human health (Ayrapetyan et al. 2015, Trends in Microbiology 23: 7). However, unlike persister cells, which quickly resume growth following drug removal, VBNC cells can remain non-growing for prolonged periods of time, thus eluding detection via traditional microbiological assays. Therefore, understanding the molecular mechanisms underlying the formation of VBNC cells requires the characterization of the clonal population with single-cell resolution. A combination of microfluidics, time-lapse microscopy, and fluorescent reporter strains offers the perfect platform for investigating individual cells while manipulating their environment.
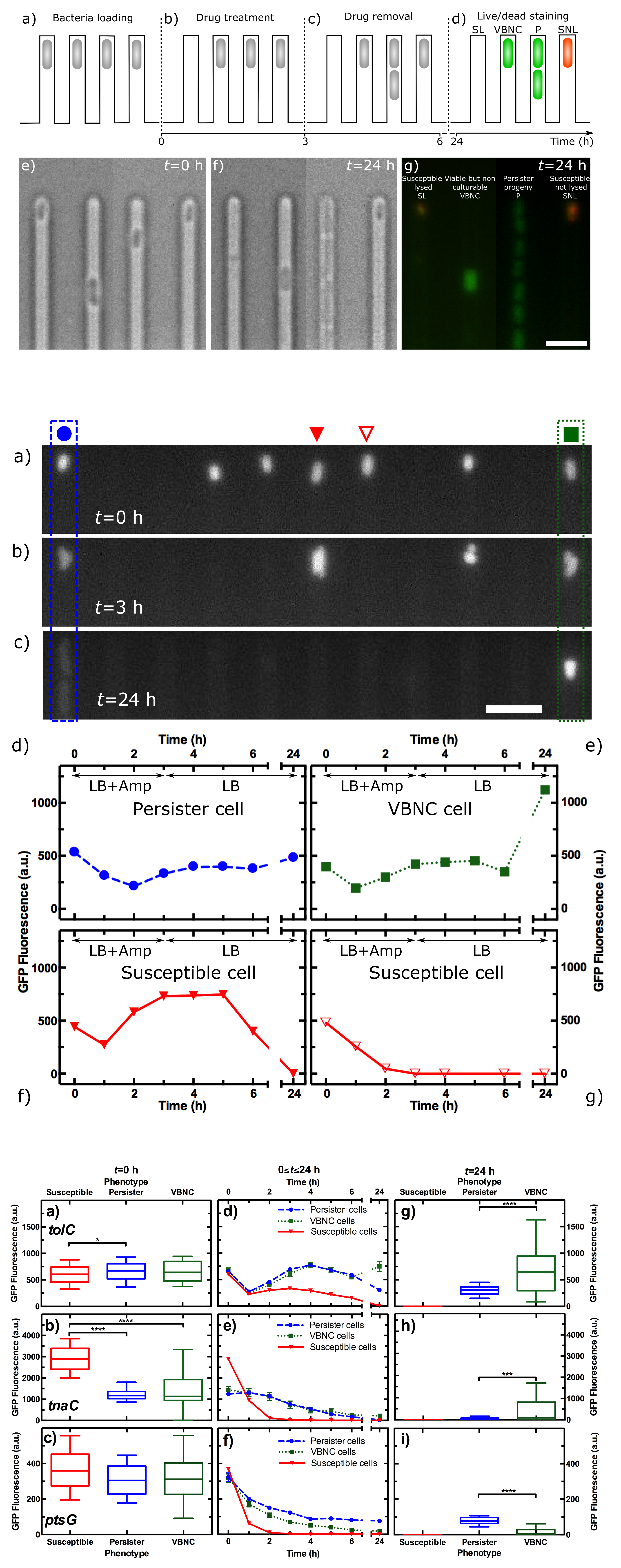
We recently developed a novel single-cell approach to investigate VBNC cells by combining microfluidics, time-lapse microscopy, and fluorescent reporter strains (Bamford et al. 2017, BMC Biology 15: 121). We perform drug treatment, bacterial culturing, and live/dead staining in series by using transcriptional reporter strains and novel adaptations to the mother machine technology. Since we track each cell throughout the experiment, we are able to quantify the size, morphology and fluorescence that each VBNC cell displayed before, during and after drug treatment.
When we used a high dose of ampicillin (25 × Minimum inhibitory concentration MIC), a β-lactam antibiotic targeting the bacterial wall, we measured that the fraction of VBNC cells (fVBNC = 0.01) was larger than the fraction of persister cells (fP =
0.003). Therefore, we provide direct evidence that VBNC cells, rather than persister cells, constitute the majority of cells surviving ampicillin treatment in a stationary phase E. coli culture.
Furthermore, we show that VBNC cells are not dead or dying cells but share similar phenotypic features with persister cells, suggesting a link between these two subpopulations, at least in the Escherichia coli strain under investigation.
We investigated reporter strains for several genes previously linked to persister and VBNC cells and found that before drug treatment, both persister and VBNC cells can be distinguished from the remainder of the population by their lower fluorescence when using a reporter strain for tnaC, encoding the leader peptide of the tnaCAB operon responsible for tryptophan metabolism. This further strengthen the link between these two antibiotic tolerant subpopulations and opens the way for a better understanding of the molecular mechanisms underlying their functioning.
Our data demonstrates the suitability of our approach for studying the physiology of non-growing
cells in response to external perturbations. Our approach will allow the identification of novel biomarkers for the isolation of VBNC and persister cells and will open new opportunities to map the detailed biochemical makeup of these clonal subpopulations.
News
- 8-September-2023: Congratulations Erin! New paper out at ISME Communications
- 10-April-2023: Congratulations Ula! New paper out at PLoS Biology
- 7-June-2022: Congratulations Ula! New paper out at eLife

